Train Models on Large Datasets
Live Notebook
You can run this notebook in a live session 
Train Models on Large Datasets¶
Most estimators in scikit-learn are designed to work with NumPy arrays or scipy sparse matricies. These data structures must fit in the RAM on a single machine.
Estimators implemented in Dask-ML work well with Dask Arrays and DataFrames. This can be much larger than a single machine’s RAM. They can be distributed in memory on a cluster of machines.
[1]:
%matplotlib inline
[2]:
from dask.distributed import Client
# Scale up: connect to your own cluster with more resources
# see http://dask.pydata.org/en/latest/setup.html
client = Client(processes=False, threads_per_worker=4,
n_workers=1, memory_limit='2GB')
client
[2]:
Client
Client-a42f0f17-0de1-11ed-a66b-000d3a8f7959
| Connection method: Cluster object | Cluster type: distributed.LocalCluster |
| Dashboard: http://10.1.1.64:8787/status |
Cluster Info
LocalCluster
770098da
| Dashboard: http://10.1.1.64:8787/status | Workers: 1 |
| Total threads: 4 | Total memory: 1.86 GiB |
| Status: running | Using processes: False |
Scheduler Info
Scheduler
Scheduler-09a424d6-c74b-456d-8b3c-65109da5de8c
| Comm: inproc://10.1.1.64/9835/1 | Workers: 1 |
| Dashboard: http://10.1.1.64:8787/status | Total threads: 4 |
| Started: Just now | Total memory: 1.86 GiB |
Workers
Worker: 0
| Comm: inproc://10.1.1.64/9835/4 | Total threads: 4 |
| Dashboard: http://10.1.1.64:33217/status | Memory: 1.86 GiB |
| Nanny: None | |
| Local directory: /home/runner/work/dask-examples/dask-examples/machine-learning/dask-worker-space/worker-by3a7q7t | |
[3]:
import dask_ml.datasets
import dask_ml.cluster
import matplotlib.pyplot as plt
In this example, we’ll use dask_ml.datasets.make_blobs to generate some random dask arrays.
[4]:
# Scale up: increase n_samples or n_features
X, y = dask_ml.datasets.make_blobs(n_samples=1000000,
chunks=100000,
random_state=0,
centers=3)
X = X.persist()
X
[4]:
|
We’ll use the k-means implemented in Dask-ML to cluster the points. It uses the k-means|| (read: “k-means parallel”) initialization algorithm, which scales better than k-means++. All of the computation, both during and after initialization, can be done in parallel.
[5]:
km = dask_ml.cluster.KMeans(n_clusters=3, init_max_iter=2, oversampling_factor=10)
km.fit(X)
/usr/share/miniconda3/envs/dask-examples/lib/python3.9/site-packages/dask/base.py:1283: UserWarning: Running on a single-machine scheduler when a distributed client is active might lead to unexpected results.
warnings.warn(
[5]:
KMeans(init_max_iter=2, n_clusters=3, oversampling_factor=10)
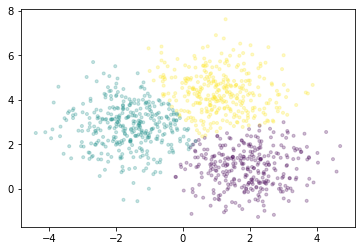
We’ll plot a sample of points, colored by the cluster each falls into.
[6]:
fig, ax = plt.subplots()
ax.scatter(X[::1000, 0], X[::1000, 1], marker='.', c=km.labels_[::1000],
cmap='viridis', alpha=0.25);
For all the estimators implemented in Dask-ML, see the API documentation.